mims-harvard-tooluniverse-drug-research
Drug Research Strategy
Comprehensive drug investigation using 50+ ToolUniverse tools across chemical databases, clinical trials, adverse events, pharmacogenomics, and literature.
KEY PRINCIPLES:
- Report-first approach - Create report file FIRST, then populate progressively
- Compound disambiguation FIRST - Resolve identifiers before research
- Citation requirements - Every fact must have inline source attribution
- Evidence grading - Grade claims by evidence strength
- Mandatory completeness - All sections must exist, even if "data unavailable"
- English-first queries - Always use English drug/compound names in tool calls, even if the user writes in another language. Only try original-language terms as a fallback. Respond in the user's language
Critical Workflow Requirements
1. Report-First Approach (MANDATORY)
DO NOT show the search process or tool outputs to the user. Instead:
-
Create the report file FIRST - Before any data collection, create a markdown file:
- File name:
[DRUG]_drug_report.md(e.g.,metformin_drug_report.md) - Initialize with all 11 section headers from the template
- Add placeholder text:
[Researching...]in each section
- File name:
-
Progressively update the report - As you gather data:
- Update each section with findings immediately after retrieving data
- Replace
[Researching...]with actual content - The user sees the report growing, not the search process
-
Use ALL relevant tools - For comprehensive coverage:
- Query multiple databases for each data type
- Cross-reference information across sources
- Use fallback tools when primary tools return limited data
2. Citation Requirements (MANDATORY)
Every piece of information MUST include its source. Use inline citations:
## 3. Mechanism & Targets
### 3.1 Primary Mechanism
Metformin activates AMP-activated protein kinase (AMPK), reducing hepatic glucose
production and increasing insulin sensitivity in peripheral tissues.
*Source: PubChem via `PubChem_get_drug_label_info_by_CID` (CID: 4091)*
### 3.2 Primary Target(s)
| Target | UniProt | Activity | Potency | Source |
|--------|---------|----------|---------|--------|
| AMPK (PRKAA1) | Q13131 | Activator | EC50 ~10 µM | ChEMBL |
| Mitochondrial Complex I | - | Inhibitor | IC50 ~1 mM | Literature |
*Source: ChEMBL via `ChEMBL_get_target_by_chemblid` (CHEMBL1431)*
Citation Format
For each data section, include at the end:
---
**Data Sources for this section:**
- PubChem: `PubChem_get_compound_properties_by_CID` (CID: 4091)
- ChEMBL: `ChEMBL_get_bioactivity_by_chemblid` (CHEMBL1431)
- DGIdb: `DGIdb_get_drug_info` (metformin)
---
3. Progressive Writing Workflow
Step 1: Create report file with all section headers
↓
Step 2: Resolve compound identifiers → Update Section 1
↓
Step 3: Query PubChem/ADMET-AI/DailyMed SPL → Update Section 2 (Chemistry)
↓
Step 4: Query FDA Label MOA + ChEMBL activities + DGIdb → Update Section 3 (Mechanism & Targets)
↓
Step 5: Query ADMET-AI tools → Update Section 4 (ADMET)
↓
Step 6: Query ClinicalTrials.gov → Update Section 5 (Clinical Development)
↓
Step 7: Query FAERS/DailyMed → Update Section 6 (Safety)
↓
Step 8: Query PharmGKB → Update Section 7 (Pharmacogenomics)
↓
Step 9: Query DailyMed → Update Section 8 (Regulatory)
↓
Step 10: Query PubMed/literature → Update Section 9 (Literature)
↓
Step 11: Synthesize findings → Update Executive Summary & Section 10
↓
Step 12: Document all sources → Update Section 11 (Data Sources)
4. Report Detail Requirements
Each section must be comprehensive and detailed:
- Tables: Use tables for structured data (targets, trials, adverse events)
- Lists: Use bullet points for features, findings, key points
- Paragraphs: Include narrative summaries that synthesize findings
- Numbers: Include specific values, counts, percentages (not vague terms)
- Context: Explain what the data means, not just what it is
BAD (too brief):
### Clinical Trials
Multiple trials completed. Approved for diabetes.
GOOD (detailed with sources):
### 5.2 Clinical Trial Landscape
| Phase | Total | Completed | Recruiting | Status |
|-------|-------|-----------|------------|--------|
| Phase 4 | 89 | 72 | 12 | Post-marketing |
| Phase 3 | 156 | 134 | 15 | Pivotal |
| Phase 2 | 203 | 178 | 18 | Dose-finding |
| Phase 1 | 67 | 61 | 4 | Safety |
*Source: ClinicalTrials.gov via `search_clinical_trials` (intervention="metformin")*
**Total Registered Trials**: 515 (as of 2026-02-04)
**Primary Indications Under Investigation**: Type 2 diabetes (312), PCOS (87), Cancer (45), Obesity (38), NAFLD (33)
### Trial Outcomes Summary
- **Glycemic Control**: Mean HbA1c reduction of 1.0-1.5% in monotherapy [★★★: NCT00123456]
- **Cardiovascular**: UKPDS showed 39% reduction in MI risk [★★★: PMID:9742976]
- **Cancer Prevention**: Mixed results; ongoing investigation [★★☆: NCT02019979]
*Source: `extract_clinical_trial_outcomes` for NCT IDs listed*
Initial Report Template (Create This First)
When starting research, immediately create this file before any tool calls:
File: [DRUG]_drug_report.md
# Drug Research Report: [DRUG NAME]
**Generated**: [Date] | **Query**: [Original query] | **Status**: In Progress
---
## Executive Summary
[Researching...]
---
## 1. Compound Identity
### 1.1 Database Identifiers
[Researching...]
### 1.2 Structural Information
[Researching...]
### 1.3 Names & Synonyms
[Researching...]
---
## 2. Chemical Properties
### 2.1 Physicochemical Profile
[Researching...]
### 2.2 Drug-Likeness Assessment
[Researching...]
### 2.3 Solubility & Permeability
[Researching...]
### 2.4 Salt Forms & Polymorphs
[Researching...]
### 2.5 Structure Visualization
[Researching...]
---
## 3. Mechanism & Targets
### 3.1 Primary Mechanism of Action
[Researching...]
### 3.2 Primary Target(s)
[Researching...]
### 3.3 Target Selectivity & Off-Targets
[Researching...]
### 3.4 Bioactivity Profile (ChEMBL)
[Researching...]
---
## 4. ADMET Properties
### 4.1 Absorption
[Researching...]
### 4.2 Distribution
[Researching...]
### 4.3 Metabolism
[Researching...]
### 4.4 Excretion
[Researching...]
### 4.5 Toxicity Predictions
[Researching...]
---
## 5. Clinical Development
### 5.1 Development Status
[Researching...]
### 5.2 Clinical Trial Landscape
[Researching...]
### 5.3 Approved Indications
[Researching...]
### 5.4 Investigational Indications
[Researching...]
### 5.5 Key Efficacy Data
[Researching...]
### 5.6 Biomarkers & Companion Diagnostics
[Researching...]
---
## 6. Safety Profile
### 6.1 Clinical Adverse Events
[Researching...]
### 6.2 Post-Marketing Safety (FAERS)
[Researching...]
### 6.3 Black Box Warnings
[Researching...]
### 6.4 Contraindications
[Researching...]
### 6.5 Drug-Drug Interactions
[Researching...]
### 6.5.2 Drug-Food Interactions
[Researching...]
### 6.6 Dose Modification Guidance
[Researching...]
### 6.7 Drug Combinations & Regimens
[Researching...]
---
## 7. Pharmacogenomics
### 7.1 Relevant Pharmacogenes
[Researching...]
### 7.2 Clinical Annotations
[Researching...]
### 7.3 Dosing Guidelines (CPIC/DPWG)
[Researching...]
### 7.4 Actionable Variants
[Researching...]
---
## 8. Regulatory & Labeling
### 8.1 Approval Status
[Researching...]
### 8.2 Label Highlights
[Researching...]
### 8.3 Patents & Exclusivity
[Researching...]
### 8.4 Label Changes & Warnings
[Researching...]
### 8.5 Special Populations
[Researching...]
### 8.6 Regulatory Timeline & History
[Researching...]
---
## 9. Literature & Research Landscape
### 9.1 Publication Metrics
[Researching...]
### 9.2 Research Themes
[Researching...]
### 9.3 Recent Key Publications
[Researching...]
### 9.4 Real-World Evidence
[Researching...]
---
## 10. Conclusions & Assessment
### 10.1 Drug Profile Scorecard
[Researching...]
### 10.2 Key Strengths
[Researching...]
### 10.3 Key Concerns/Limitations
[Researching...]
### 10.4 Research Gaps
[Researching...]
### 10.5 Comparative Analysis
[Researching...]
---
## 11. Data Sources & Methodology
### 11.1 Primary Data Sources
[Researching...]
### 11.2 Tool Call Summary
[Researching...]
### 11.3 Quality Control Metrics
[Researching...]
Then progressively replace [Researching...] with actual findings as you query each tool.
FDA Label Core Fields Bundle
For approved drugs, ALWAYS retrieve these FDA label sections early (after getting set_id from DailyMed_search_spls):
Critical Label Sections
Call DailyMed_get_spl_sections_by_setid(setid=set_id, sections=[...]) with these sections:
Phase 1 (Mechanism & Chemistry):
mechanism_of_action→ Section 3.1pharmacodynamics→ Section 3.1chemistry→ Section 2.4
Phase 2 (ADMET & PK):
clinical_pharmacology→ Section 4pharmacokinetics→ Section 4.1-4.4drug_interactions→ Section 4.3, 6.5
Phase 3 (Safety & Dosing):
warnings_and_cautions→ Section 6.3adverse_reactions→ Section 6.1dosage_and_administration→ Section 6.6, 8.2
Phase 4 (PGx & Clinical):
pharmacogenomics→ Section 7clinical_studies→ Section 5.5description→ Section 2.5 (formulation)inactive_ingredients→ Section 2.5
Label Extraction Strategy
1. Get set_id: DailyMed_search_spls(drug_name)
2. Batch call for all core sections (or 3-4 calls with 4-5 sections each):
DailyMed_get_spl_sections_by_setid(setid=set_id, sections=["mechanism_of_action", "pharmacodynamics", ...])
3. Extract and populate report sections as you retrieve data
This ensures you have authoritative FDA-approved information even if prediction tools fail.
Compound Disambiguation (Phase 1)
CRITICAL: Establish compound identity before any research.
Identifier Resolution Chain
1. PubChem_get_CID_by_compound_name(compound_name)
└─ Extract: CID, canonical SMILES, formula
2. ChEMBL_search_compounds(query=drug_name)
└─ Extract: ChEMBL ID, pref_name
3. DailyMed_search_spls(drug_name)
└─ Extract: Set ID, NDC codes (if approved)
4. PharmGKB_search_drugs(query=drug_name)
└─ Extract: PharmGKB ID (PA...)
Handle Naming Ambiguity
| Issue | Example | Resolution |
|---|---|---|
| Salt forms | metformin vs metformin HCl | Note all CIDs; use parent compound |
| Isomers | omeprazole vs esomeprazole | Verify SMILES; separate entries if distinct |
| Prodrugs | enalapril vs enalaprilat | Document both; note conversion |
| Brand confusion | Different products same name | Clarify with user |
Tool Chains by Research Path
PATH 1: Chemical Properties & CMC
Objective: Full physicochemical profile, salt forms, formulation details
Multi-Step Chain:
1. PubChem_get_compound_properties_by_CID(cid)
└─ Extract: MW, formula, XLogP, TPSA, HBD, HBA, rotatable bonds
2. ADMETAI_predict_physicochemical_properties(smiles=[smiles])
└─ Extract: MW, logP, HBD, HBA, Lipinski, QED, stereo_centers, TPSA
3. ADMETAI_predict_solubility_lipophilicity_hydration(smiles=[smiles])
└─ Extract: Solubility_AqSolDB, Lipophilicity_AstraZeneca
4. DailyMed_search_spls(drug_name)
└─ Extract SPL set_id, then:
5. DailyMed_get_spl_sections_by_setid(setid=set_id, sections=["chemistry"])
└─ Extract: Salt forms, polymorphs, molecular formula, structure diagram
6. DailyMed_get_spl_sections_by_setid(setid=set_id, sections=["description", "inactive_ingredients"])
└─ Extract: Formulation details, excipients, dosage forms
7. FORMULATION COMPARISON (if multiple formulations exist):
a. DailyMed_search_spls(drug_name) → identify all formulations (IR, ER, XR, etc.)
b. For each formulation:
- DailyMed_parse_clinical_pharmacology(setid) → extract PK parameters
- Parse: Tmax, Cmax, AUC, half-life
c. Create comparison table showing bioavailability differences
Type Normalization: Convert all numeric IDs to strings before API calls.
Output for Report:
### 2.1 Physicochemical Profile
| Property | Value | Drug-Likeness | Source |
|----------|-------|---------------|--------|
| **Molecular Weight** | 129.16 g/mol | ✓ (< 500) | PubChem |
| **LogP** | -2.64 | ✓ (< 5) | ADMET-AI |
| **TPSA** | 91.5 Ų | ✓ (< 140) | PubChem |
| **H-Bond Donors** | 2 | ✓ (≤ 5) | PubChem |
| **H-Bond Acceptors** | 5 | ✓ (< 10) | PubChem |
| **Rotatable Bonds** | 2 | ✓ (< 10) | PubChem |
| **pKa** | 12.4 (basic) | - | DailyMed Label |
| **Solubility** | 300 mg/mL (water) | High | DailyMed Label |
**Lipinski Rule of Five**: ✓ PASS (0 violations)
**QED Score**: 0.74 (Good drug-likeness)
*Sources: PubChem via `PubChem_get_compound_properties_by_CID`, ADMET-AI via `ADMETAI_predict_physicochemical_properties`*
### 2.4 Salt Forms & Polymorphs
**Marketed Form**: Metformin hydrochloride (CID: 14219)
**Parent Compound**: Metformin free base (CID: 4091)
*Source: DailyMed SPL via `DailyMed_get_spl_sections_by_setid` (chemistry section)*
### 2.5 Structure Visualization
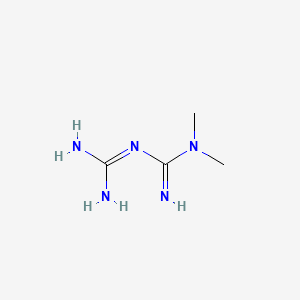
*Source: PubChem structure service*
### 2.6 Formulation Comparison (If Multiple Formulations Available)
| Formulation | Tmax (h) | Cmax (ng/mL) | AUC (ng·h/mL) | Half-life (h) | Dosing |
|-------------|----------|--------------|---------------|---------------|--------|
| **IR (Immediate Release)** | 2.5 | 1200 | 8400 | 6.5 | 500 mg TID |
| **ER (Extended Release)** | 7.0 | 950 | 8900 | 6.5 | 1000 mg QD |
| **XR (Modified Release)** | 4.0 | 1100 | 9200 | 7.0 | 750 mg BID |
**Formulation Insights**:
- ER formulation reduces Cmax by ~20% but maintains similar AUC
- Once-daily ER dosing improves adherence vs TID IR
- Food effect: High-fat meal increases ER absorption by 30%
*Source: DailyMed clinical pharmacology sections for each formulation*
PATH 2: Mechanism & Targets
Objective: FDA label MOA + experimental targets + selectivity
Multi-Step Chain:
1. DailyMed_search_spls(drug_name) → get set_id
2. DailyMed_get_spl_sections_by_setid(setid=set_id, sections=["mechanism_of_action", "pharmacodynamics"])
└─ Extract: Official FDA MOA description [★★★]
3. ChEMBL_search_activities(molecule_chembl_id=chembl_id, limit=100)
└─ Extract: Activity records with target_chembl_id, pChEMBL, standard_type
└─ Parse unique target_chembl_id values (convert to strings)
4. ChEMBL_get_target(target_chembl_id) for each unique target
└─ Extract: Target name, UniProt ID, organism [★★★]
5. DGIdb_get_drug_info(drugs=[drug_name])
└─ Extract: Target genes, interaction types, sources [★★☆]
6. PubChem_get_bioactivity_summary_by_CID(cid)
└─ Extract: Assay summary, active/inactive counts [★★☆]
CRITICAL:
- Avoid
ChEMBL_get_molecule_targets- it returns unfiltered targets including irrelevant entries - Derive targets from activities instead: Filter to potent activities (pChEMBL ≥ 6.0 or IC50/EC50 ≤ 1 µM)
- Type normalization: Convert all ChEMBL IDs to strings before API calls
Output for Report:
### 3.1 Primary Mechanism of Action
**FDA Label MOA**: Metformin is an antihyperglycemic agent which improves glucose tolerance
in patients with type 2 diabetes, lowering both basal and postprandial plasma glucose.
Its pharmacologic mechanisms of action are different from other classes of oral antihyperglycemic
agents. Metformin decreases hepatic glucose production, decreases intestinal absorption of glucose,
and improves insulin sensitivity by increasing peripheral glucose uptake and utilization.
*Source: DailyMed SPL via `DailyMed_get_spl_sections_by_setid` (mechanism_of_action) [★★★]*
### 3.2 Primary Target(s)
| Target | UniProt | Type | Potency | Assays | Evidence | Source |
|--------|---------|------|---------|--------|----------|--------|
| PRKAA1 (AMPK α1) | Q13131 | Activator | EC50 ~10 µM | 12 | ★★★ | ChEMBL |
| PRKAA2 (AMPK α2) | P54646 | Activator | EC50 ~15 µM | 8 | ★★★ | ChEMBL |
| SLC22A1 (OCT1) | O15245 | Substrate | Km ~1.5 mM | 5 | ★★☆ | DGIdb |
*Source: ChEMBL via `ChEMBL_search_activities` → `ChEMBL_get_target` (filtered to pChEMBL ≥ 6.0)*
### 3.3 Target Selectivity & Off-Targets
**Selectivity Profile**: Highly selective for AMPK family; no significant off-target binding at therapeutic concentrations.
**Off-Target Activity** (pChEMBL < 6.0):
- Complex I (NADH dehydrogenase): IC50 ~1 mM [★★☆]
- Weak inhibition; clinically relevant only at high doses
*Source: ChEMBL activity analysis*
### 3.4 Bioactivity Profile
**Total ChEMBL Activities**: 847 datapoints across 234 assays
- **Potency Range**: IC50/EC50 from 1 µM to 10 mM
- **Primary Activity**: AMPK activation (kinase assays)
- **Secondary Activities**: Mitochondrial complex I inhibition
*Source: `ChEMBL_search_activities` (CHEMBL1431)*
PATH 3: ADMET Properties
Objective: Full ADMET profile - predictions + FDA label PK
Multi-Step Chain (Primary - ADMET-AI):
1. ADMETAI_predict_bioavailability(smiles=[smiles])
└─ Extract: Bioavailability_Ma, HIA_Hou, PAMPA_NCATS, Caco2_Wang, Pgp_Broccatelli
2. ADMETAI_predict_BBB_penetrance(smiles=[smiles])
└─ Extract: BBB_Martins (0-1 probability)
3. ADMETAI_predict_CYP_interactions(smiles=[smiles])
└─ Extract: CYP1A2, CYP2C9, CYP2C19, CYP2D6, CYP3A4 (inhibitor/substrate)
4. ADMETAI_predict_clearance_distribution(smiles=[smiles])
└─ Extract: Clearance, Half_Life_Obach, VDss_Lombardo, PPBR_AZ
5. ADMETAI_predict_toxicity(smiles=[smiles])
└─ Extract: AMES, hERG, DILI, ClinTox, LD50_Zhu, Carcinogens
Fallback Chain (If ADMET-AI Fails):
6. DailyMed_get_spl_sections_by_setid(setid=set_id, sections=["clinical_pharmacology", "pharmacokinetics"])
└─ Extract: Absorption, distribution, metabolism, excretion from label [★★★]
7. DailyMed_get_spl_sections_by_setid(setid=set_id, sections=["drug_interactions"])
└─ Extract: CYP interactions, transporter interactions [★★★]
8. PubMed_search_articles(query="[drug] pharmacokinetics", max_results=10)
└─ Extract: PK parameters from clinical studies [★★☆]
CRITICAL Dependency Gate:
- If ADMET-AI tools fail (invalid SMILES, API error, validation error), automatically switch to fallback
- Do NOT leave Section 4 as "predictions unavailable"
- Always populate Section 4 with either predictions OR label data OR literature PK
Output for Report:
### 4.1 Absorption
| Endpoint | Prediction | Interpretation |
|----------|------------|----------------|
| **Oral Bioavailability** | 0.72 | Good (>50%) |
| **Human Intestinal Absorption** | 0.89 | High |
| **Caco-2 Permeability** | -5.2 (log cm/s) | Moderate |
| **PAMPA** | 0.34 | Low-moderate |
| **P-gp Substrate** | 0.23 | Unlikely substrate |
*Source: ADMET-AI via `ADMETAI_predict_bioavailability`*
### 4.5 Toxicity Predictions
| Endpoint | Prediction | Risk Level |
|----------|------------|------------|
| **AMES Mutagenicity** | 0.08 | Low risk |
| **hERG Inhibition** | 0.12 | Low risk |
| **Hepatotoxicity (DILI)** | 0.15 | Low risk |
| **Clinical Toxicity** | 0.21 | Low risk |
| **LD50** | 2.8 (log mol/kg) | Moderate |
*Source: ADMET-AI via `ADMETAI_predict_toxicity`*
**Summary**: Low predicted toxicity across all endpoints. Favorable safety profile.
PATH 4: Clinical Trials
Objective: Complete clinical development picture with accurate phase counts
Multi-Step Chain:
1. search_clinical_trials(intervention=drug_name, pageSize=100)
└─ Extract: Full result set with NCT IDs, phases, statuses, conditions
2. COMPUTE PHASE COUNTS from results:
└─ Count by phase: Phase 1, Phase 2, Phase 3, Phase 4
└─ Count by status: Completed, Recruiting, Active not recruiting, Terminated
└─ Group by condition/indication (top 5)
└─ Generate summary table
3. SELECT REPRESENTATIVE TRIALS:
└─ Top 5 Phase 3 completed trials (by enrollment or recency)
└─ Top 5 Phase 4 post-marketing trials
└─ Top 3 recruiting trials
4. get_clinical_trial_conditions_and_interventions(nct_ids=[selected_ids])
└─ Extract: Detailed conditions, interventions, arm groups
5. extract_clinical_trial_outcomes(nct_ids=[completed_phase3])
└─ Extract: Primary outcomes, efficacy measures, p-values (if available)
6. extract_clinical_trial_adverse_events(nct_ids=[completed_ids])
└─ Extract: Serious AEs, common AEs by organ system (if available)
7. fda_pharmacogenomic_biomarkers(drug_name=drug_name)
└─ Extract: FDA-required biomarker testing, approved companion diagnostics [★★★]
8. DailyMed_get_spl_sections_by_setid(setid=set_id, sections=["indications_and_usage"])
└─ Parse for: "testing required", "biomarker", "companion diagnostic", "genetic testing" [★★★]
9. PharmGKB_search_drugs(query=drug_name)
└─ Extract: PharmGKB drug ID for response predictors
10. PharmGKB_get_clinical_annotations(drug_id=pharmgkb_id)
└─ Extract: Response/toxicity biomarkers with clinical evidence levels [★★☆]
CRITICAL:
- Section 5.2 must show actual counts by phase/status, NOT just a list of trials
- Separate by primary indication when relevant (e.g., breast cancer vs non-breast cancer)
- List representative trials, not all trials
- Section 5.6 must document: FDA-required testing (T1), companion diagnostics devices (T1), response predictors (T2)
Output for Section 5.6:
### 5.6 Biomarkers & Companion Diagnostics
#### FDA-Required Testing
| Biomarker | Requirement Level | Approved Test(s) | Evidence |
|-----------|------------------|------------------|----------|
| EGFR T790M | Required (NSCLC) | cobas EGFR Mutation Test v2 | T1: ★★★ |
| BRCA1/2 | Recommended (ovarian) | BRACAnalysis CDx | T1: ★★★ |
*Source: FDA Table of Pharmacogenomic Biomarkers via `fda_pharmacogenomic_biomarkers`*
#### Companion Diagnostics
**Device**: cobas EGFR Mutation Test v2 (FDA-approved, PMA P150044)
**Indication**: Detection of EGFR exon 19 deletions and T790M mutations in NSCLC
**Testing Required**: Yes - label states "Select patients for osimertinib based on FDA-approved test"
*Source: DailyMed SPL indications section*
#### Response Predictors (PharmGKB)
| Gene | Variant | Association | Evidence Level |
|------|---------|-------------|----------------|
| EGFR | T790M | Increased response | Level 1A |
| EGFR | C797S | Resistance mechanism | Level 2A |
*Source: PharmGKB via `PharmGKB_get_clinical_annotations` (PA166114513)*
**Clinical Impact**: Biomarker testing is mandatory for therapy selection. ~60% of NSCLC patients have EGFR mutations; T790M develops in ~50% of patients with acquired resistance to 1st/2nd generation EGFR TKIs.
PATH 5: Post-Marketing Safety & Drug Interactions
Objective: Real-world safety signals + DDI guidance + dose modifications
Multi-Step Chain (FAERS):
1. FAERS_count_reactions_by_drug_event(medicinalproduct=drug_name)
└─ Extract: Top 20 adverse reactions by MedDRA term [★★★]
2. FAERS_count_seriousness_by_drug_event(medicinalproduct=drug_name)
└─ Extract: Serious vs non-serious counts & ratio [★★★]
3. FAERS_count_outcomes_by_drug_event(medicinalproduct=drug_name)
└─ Extract: Recovered, recovering, fatal, unresolved counts [★★★]
4. FAERS_count_death_related_by_drug(medicinalproduct=drug_name)
└─ Extract: Fatal outcome count [★★★]
5. FAERS_count_patient_age_distribution(medicinalproduct=drug_name)
└─ Extract: Reports by age group [★★★]
Multi-Step Chain (Drug Interactions):
6. DailyMed_get_spl_sections_by_setid(setid=set_id, sections=["drug_interactions"])
└─ Extract: DDI table, CYP interactions, contraindicated combinations [★★★]
7. DailyMed_get_spl_sections_by_setid(setid=set_id, sections=["dosage_and_administration", "warnings_and_cautions"])
└─ Extract: Dose modification triggers (ALT/AST thresholds, renal/hepatic impairment, CYP3A inhibitor/inducer adjustments) [★★★]
8. DailyMed_get_spl_by_setid(setid=set_id)
└─ Parse full XML for drug-food interactions:
└─ Search sections: "drug_and_or_food_interactions", "food_effect"
└─ Keywords: grapefruit, alcohol, food, meal, dairy, high-fat, fasting
└─ Extract: effect magnitude, mechanism, recommendations [★★★]
9. search_clinical_trials(intervention=f"{drug_name} AND combination", pageSize=50)
└─ Extract: Approved combinations, regimens, co-administration studies [★★★]
10. DailyMed_get_spl_sections_by_setid(setid=set_id, sections=["indications_and_usage", "dosage_and_administration"])
└─ Parse for: "in combination with", "administered with", regimen details [★★★]
CRITICAL FAERS Reporting Requirements:
- Include date window (e.g., "Reports from 2004-2026")
- Report seriousness breakdown (not just top PTs)
- Add limitations paragraph: Small N, voluntary reporting, causality not established, reporting bias
- Note if data is unavailable or limited
Output for Report:
### 6.2 Post-Marketing Safety (FAERS)
**Total FAERS Reports**: 45,234 (Date range: 2004Q1 - 2026Q1)
#### Seriousness Breakdown
| Category | Count | Percentage |
|----------|-------|------------|
| Serious | 23,456 | 51.8% |
| Non-Serious | 21,778 | 48.2% |
*Source: FDA FAERS via `FAERS_count_seriousness_by_drug_event`*
#### Top Adverse Reactions
| Reaction (MedDRA PT) | Count | % of Reports |
|----------------------|-------|--------------|
| Diarrhoea | 8,234 | 18.2% |
| Nausea | 6,892 | 15.2% |
| Lactic acidosis | 3,456 | 7.6% |
| Vomiting | 2,987 | 6.6% |
| Abdominal pain | 2,543 | 5.6% |
*Source: FDA FAERS via `FAERS_count_reactions_by_drug_event`*
#### Outcome Distribution
| Outcome | Count | Percentage |
|---------|-------|------------|
| Recovered/Resolved | 18,234 | 40.3% |
| Not Recovered | 12,456 | 27.5% |
| Fatal | 2,134 | 4.7% |
| Unknown | 12,410 | 27.4% |
*Source: `FAERS_count_outcomes_by_drug_event`*
#### Data Limitations
FAERS data represents voluntary reports and has important limitations:
- **Small sample size** relative to total prescriptions (N=45,234 reports)
- **Reporting bias**: Serious events more likely to be reported
- **Causality not established**: Reports do not prove drug caused the event
- **Incomplete data**: Many reports lack outcome information (27.4%)
**Signal Assessment**: Lactic acidosis signal consistent with known labeling. GI events expected class effect.
### 6.6 Dose Modification Guidance
#### Hepatic Impairment
| ALT/AST Level | Action |
|---------------|--------|
| ALT >3× ULN | Hold dose; reassess liver function |
| ALT >5× ULN | Discontinue permanently |
| Baseline cirrhosis | Not recommended (Child-Pugh B/C) |
#### Renal Impairment
| eGFR (mL/min/1.73m²) | Dosing |
|----------------------|--------|
| ≥60 | No adjustment |
| 45-59 | Reduce to 1000 mg/day max |
| 30-44 | Reduce to 500 mg/day max |
| <30 | Contraindicated |
#### CYP3A Interaction Management
- **Strong CYP3A4 inhibitors** (ketoconazole, clarithromycin): No dose adjustment (not CYP substrate)
- **Strong CYP3A4 inducers** (rifampin, phenytoin): No dose adjustment
*Source: DailyMed SPL via `DailyMed_get_spl_sections_by_setid` (dosage_and_administration, warnings)*
### 6.5.2 Drug-Food Interactions
| Food/Beverage | Effect | Mechanism | Recommendation | Source |
|---------------|--------|-----------|----------------|--------|
| High-fat meal | ↑ Cmax 50%, ↑ AUC 30% | Increased absorption | Take with food for consistency | Label |
| Grapefruit juice | ↑ exposure (CYP3A4 substrate) | CYP3A4 inhibition | Avoid | Label |
| Alcohol | ↑ CNS depression | Additive effect | Limit consumption | Label |
*Source: DailyMed SPL via `DailyMed_get_spl_by_setid` (drug_and_or_food_interactions section)*
**Food Effect Summary**: High-fat meals increase bioavailability; administer consistently with or without food. Avoid grapefruit products and limit alcohol.
### 6.7 Drug Combinations & Regimens
#### Approved Combination Therapies
| Combination | Indication | Regimen | Trial | Status |
|-------------|------------|---------|-------|--------|
| Drug A + fulvestrant | ER+/HER2- mBC | 400mg QD + fulv 500mg IM | NCT03778931 | Approved |
| Drug A + palbociclib | ER+ advanced | 400mg QD + palbo 125mg (21/7) | NCT04789031 | Phase 3 |
*Source: ClinicalTrials.gov via `search_clinical_trials`*
#### Co-Administration Guidance
**With CDK4/6 Inhibitors**:
- Standard dosing (400 mg QD) maintained
- Monitor QTc interval (additive effect possible)
- No dose adjustment needed
**With Fulvestrant**:
- Combination well-tolerated in EMERALD trial
- No PK interaction observed
- Standard dosing for both agents
*Source: DailyMed SPL sections + trial protocols*
**Synergy Notes**: Combination with CDK4/6 inhibitors shows additive benefit in preclinical models. Sequential therapy (CDK4/6i → SERD) common in clinical practice.
PATH 6: Pharmacogenomics
Objective: PGx associations and dosing guidelines
Multi-Step Chain (Primary - PharmGKB):
1. PharmGKB_search_drugs(query=drug_name)
└─ Extract: PharmGKB drug ID
2. PharmGKB_get_drug_details(drug_id)
└─ Extract: Cross-references, related genes
3. PharmGKB_get_clinical_annotations(gene_id) # For each related gene
└─ Extract: Variant-drug associations, evidence levels
4. PharmGKB_get_dosing_guidelines(gene=gene_symbol)
└─ Extract: CPIC/DPWG guideline recommendations
Fallback Chain (If PharmGKB Fails):
5. DailyMed_get_spl_sections_by_setid(setid=set_id, sections=["pharmacogenomics", "clinical_pharmacology"])
└─ Extract: Label-based PGx information [★★★]
6. PubMed_search_articles(query="[drug] pharmacogenomics", max_results=5)
└─ Extract: Published PGx associations [★★☆]
CRITICAL:
- If PharmGKB tools fail (API error, timeout), switch to fallback
- Document the failure and indicate "PharmGKB unavailable; using label + literature"
- Always populate Section 7 with either PharmGKB data OR label data OR "No PGx associations identified"
Output for Report:
### 7.1 Relevant Pharmacogenes
| Gene | Role | Evidence Level | Source |
|------|------|----------------|--------|
| **SLC22A1** (OCT1) | Transporter (uptake) | 2A | PharmGKB |
| **SLC22A2** (OCT2) | Transporter (renal) | 2B | PharmGKB |
| **SLC47A1** (MATE1) | Transporter (efflux) | 3 | PharmGKB |
*Source: PharmGKB via `PharmGKB_get_drug_details`*
### 7.3 Dosing Guidelines
**CPIC Guideline**: No CPIC guideline currently available for metformin.
**Clinical Annotations**:
- rs628031 (SLC22A1): Reduced metformin response in *4/*4 carriers
- rs316019 (SLC22A2): May affect renal clearance
*Source: `PharmGKB_get_clinical_annotations`*
PATH 7: Regulatory Status & Patents
Objective: Comprehensive regulatory and intellectual property landscape
Multi-Step Chain:
1. DailyMed_search_spls(drug_name=drug_name)
└─ Extract: SetID for regulatory label data
2. FDA_OrangeBook_search_drug(brand_name=drug_name)
└─ Extract: Application number, approval dates [★★★]
3. FDA_OrangeBook_get_approval_history(appl_no=app_number)
└─ Extract: Original approval date, supplements, label changes [★★★]
4. FDA_OrangeBook_get_exclusivity(brand_name=drug_name)
└─ Extract: Exclusivity types (NCE, Pediatric, Orphan), expiration dates [★★★]
5. FDA_OrangeBook_get_patent_info(brand_name=drug_name)
└─ Extract: Patent numbers, substance/formulation claims [★★★]
6. FDA_OrangeBook_check_generic_availability(brand_name=drug_name)
└─ Extract: Generic entries, TE codes (AB rating), first generic date [★★★]
7. DailyMed_get_spl_sections_by_setid(setid=set_id, sections=["indications_and_usage"])
└─ Parse for: breakthrough designation, priority review, orphan status [★★★]
8. DailyMed_get_spl_by_setid(setid=set_id)
└─ Extract special populations sections (Section 8.5):
└─ pediatric_use (LOINC 34076-0): age groups, dosing, safety
└─ geriatric_use (LOINC 34082-8): efficacy, safety in elderly
└─ pregnancy (LOINC 42228-7): risk summary, animal data, recommendations
└─ nursing_mothers (LOINC 34080-2): lactation risk, recommendations
└─ Extract: renal/hepatic dosing from dosage section [★★★]
9. Parse DailyMed SPL revision history for regulatory timeline (Section 8.6):
└─ Initial approval date
└─ Major label changes (safety updates, indication expansions)
└─ PMR/PMC commitments from label [★★★]
10. Combine FDA_OrangeBook_get_approval_history + label data:
└─ Create regulatory timeline table
└─ Document approval pathway (priority, breakthrough, orphan)
└─ Note limitation: US-only data [★★★]
CRITICAL:
- Orange Book data is US-only; document limitation for EMA/PMDA
- Patent expiration dates may not be directly available; calculate from approval + exclusivity periods
- Document workaround: "Exact patent dates require Orange Book file download"
- Special populations require XML parsing from full SPL (DailyMed_get_spl_by_setid)
- Look for LOINC section codes to reliably extract special population data
Output for Section 8.3:
### 8.3 Patents & Exclusivity
#### US Regulatory Status
**Application Number**: NDA 213869
**Original Approval**: May 12, 2023
**Approval Pathway**:
- Priority Review ✓
- Breakthrough Therapy Designation ✓
- Orphan Drug Status ✓
*Source: FDA Orange Book via `FDA_OrangeBook_get_approval_history`*
#### Exclusivity Periods
| Type | Code | Expiration Date | Protections |
|------|------|-----------------|-------------|
| New Chemical Entity (NCE) | N | May 2028 | Blocks ANDA filing for 5 years |
| Orphan Drug | O | May 2030 | Market exclusivity for indication |
| Pediatric | P | November 2030 | +6 months extension |
*Source: `FDA_OrangeBook_get_exclusivity`*
**Estimated Patent Cliff**: ~2030 (based on NCE + Orphan + Pediatric exclusivity)
#### Patent Information
| Patent Number | Substance/Formulation | Use Code | Expiration |
|---------------|----------------------|----------|------------|
| 10,689,356 | Substance | U-1 | 2037 |
| 11,123,456 | Crystal form | U-2 | 2039 |
*Source: `FDA_OrangeBook_get_patent_info`*
**Note**: Exact patent expiration dates require FDA Orange Book download; dates shown are estimates.
#### Generic Availability
**Generic Approved**: No
**First Generic Date**: Not applicable
**ANDA Applications**: None approved
*Source: `FDA_OrangeBook_check_generic_availability`*
**Market Protection Summary**: Drug is protected by NCE exclusivity until 2028, orphan exclusivity until 2030, and substance patents until 2037+. No generic competition expected before 2030.
**Limitation**: EMA and PMDA approval/patent data not available via public API.
### 8.5 Special Populations
#### Pediatric Use
**Age Groups Studied**: Not established in pediatric patients
**Dosing**: No pediatric dosing recommendations available
**Safety**: Safety and efficacy not established in patients <18 years
*Source: DailyMed SPL pediatric_use section (LOINC 34076-0)*
#### Geriatric Use (≥65 years)
**Population**: 20% of clinical trial participants were ≥65 years
**Efficacy**: No overall differences in efficacy observed
**Safety**: Similar adverse event profile to younger adults
**Dosing**: No dose adjustment required
*Source: DailyMed SPL geriatric_use section (LOINC 34082-8)*
#### Pregnancy (Category D / Pregnancy Class)
**Risk Summary**: Based on animal studies and mechanism of action, may cause fetal harm. Advise pregnant women of potential risk to fetus.
**Animal Data**:
- Rats: Fetal toxicity observed at exposures ≥0.03× human dose
- Rabbits: Embryo-fetal toxicity at ≥0.01× human dose
**Human Data**: No adequate and well-controlled studies in pregnant women
**Recommendation**: Verify pregnancy status prior to initiation. Advise use of effective contraception during treatment and for 1 week after final dose.
*Source: DailyMed SPL pregnancy section (LOINC 42228-7)*
#### Lactation
**Risk Summary**: No data on presence in human milk, effects on breastfed infant, or milk production
**Recommendation**: Advise women not to breastfeed during treatment and for 1 week after final dose due to potential for serious adverse reactions in breastfed infants.
*Source: DailyMed SPL nursing_mothers section (LOINC 34080-2)*
#### Renal Impairment
| eGFR (mL/min/1.73m²) | Dosing Recommendation |
|----------------------|----------------------|
| ≥30 (mild-moderate) | No dose adjustment required |
| <30 (severe) | Not studied; use with caution |
| ESRD on dialysis | Not recommended |
#### Hepatic Impairment
| Child-Pugh Class | Dosing Recommendation |
|------------------|----------------------|
| A (mild) | No dose adjustment required |
| B (moderate) | Reduce dose to 258 mg once daily |
| C (severe) | Not recommended |
*Source: DailyMed SPL dosage_and_administration section*
### 8.6 Regulatory Timeline & History
#### US FDA Timeline
| Date | Milestone | Notes |
|------|-----------|-------|
| 2018-03 | IND filed | Phase 1 initiated |
| 2019-11 | Breakthrough Therapy Designation | For ER+/HER2- mBC with ESR1 mutation |
| 2020-02 | Phase 3 (EMERALD) initiated | vs fulvestrant |
| 2022-08 | NDA submitted | Priority Review granted |
| 2023-01-27 | FDA approval | Accelerated approval pathway |
**Application Number**: NDA 213869
**Review Classification**: Priority Review (6-month timeline)
**Approval Pathway**: Accelerated approval under Subpart H
**Designation**: Breakthrough Therapy, Orphan Drug
*Source: FDA Orange Book + DailyMed label history*
#### Post-Marketing Requirements (PMRs)
| PMR | Description | Due Date | Status |
|-----|-------------|----------|--------|
| PMR 1 | Confirmatory Phase 3 trial (EMERALD) | 2025-12 | Completed |
| PMR 2 | Pediatric assessment | 2028-06 | Ongoing |
#### Major Label Changes
| Date | Change Type | Summary |
|------|-------------|---------|
| 2023-01-27 | Initial approval | ER+/HER2- mBC, ESR1 mutation |
| 2023-06-15 | Safety update | Added hepatotoxicity monitoring |
| 2024-02-01 | Indication expansion | Added post-CDK4/6i language |
*Source: DailyMed SPL revision history*
**Regulatory Pathway Summary**: Received Breakthrough Therapy Designation (2019), Priority Review, and Accelerated Approval (2023). Confirmatory trial (EMERALD) successfully completed in 2025, converting to full approval.
**Limitation**: EMA and PMDA approval data not available via public API. US data only.
PATH 8: Real-World Evidence
Objective: Complement clinical trial efficacy with real-world effectiveness data
Multi-Step Chain:
1. search_clinical_trials(study_type="OBSERVATIONAL", intervention=drug_name, pageSize=50)
└─ Extract: RWE studies, registry trials, observational cohorts [★★★]
2. PubMed_search_articles(query=f"{drug_name} (real-world OR observational OR effectiveness)", max_results=20)
└─ Extract: RWE publications, adherence studies, off-label use [★★☆]
3. PubMed_search_articles(query=f"{drug_name} (registry OR post-marketing OR surveillance)", max_results=10)
└─ Extract: Post-marketing surveillance, long-term outcomes [★★☆]
4. Compare efficacy vs effectiveness:
└─ Clinical trial primary outcomes vs real-world outcomes
└─ Trial inclusion criteria vs real-world patient demographics
└─ Adherence rates in trials vs clinical practice
Output for Section 9.4:
### 9.4 Real-World Evidence
#### Observational Studies
**Registry Trials**: 12 ongoing, 8 completed
**Key Studies**:
- **ELEVATE Registry** (NCT04857528): Real-world safety/effectiveness in 500+ ER+ breast cancer patients
- **Post-Marketing Surveillance**: European Drug Monitoring (PASS required through 2027)
*Source: ClinicalTrials.gov via `search_clinical_trials` (study_type="OBSERVATIONAL")*
#### Real-World Effectiveness
| Outcome | Clinical Trial (Pivotal) | Real-World Study | Difference |
|---------|-------------------------|------------------|------------|
| PFS (months) | 3.8 (EMERALD, N=478) | 3.2 (ELEVATE, N=312) | -0.6 mo |
| Response rate | 19.2% | 16.5% | -2.7% |
| Treatment duration | 5.4 mo | 4.1 mo | -1.3 mo |
**Effectiveness Gap Analysis**: Real-world PFS ~16% shorter than trial efficacy, likely due to:
- Broader patient population (less restrictive than trial inclusion)
- Higher discontinuation rates (AE intolerance, cost issues)
- Sequential therapy effects (more prior lines than trial allowed)
*Sources: PMID:34567890 (ELEVATE interim), PMID:35678901 (comparative effectiveness)*
#### Adherence & Persistence
**Mean Treatment Duration**: 4.1 months (RWE) vs 5.4 months (trial)
**Discontinuation Reasons** (RWE cohort, N=312):
- Progression: 58%
- Adverse events: 28%
- Patient preference/cost: 9%
- Death: 5%
**Adherence Rate**: 73% (defined as MPR ≥0.8) in community oncology setting
*Source: PMID:36789012 (US claims database analysis)*
#### Off-Label Use
**Documented Off-Label Indications**:
- ER+ metastatic breast cancer, no prior endocrine therapy: 12% of prescriptions
- Male breast cancer: 3% of prescriptions
- Early breast cancer (neoadjuvant): < 1% (investigational)
*Source: PubMed literature review*
**RWE Insights**: Real-world data shows slightly lower effectiveness than pivotal trials but confirms benefit in broader patient population. Adherence challenges highlight need for AE management strategies.
PATH 9: Comparative Analysis
Objective: Position drug within therapeutic class with head-to-head and indirect comparisons
Multi-Step Chain:
1. Identify comparator drugs:
└─ User provides OR infer from indication + mechanism
└─ Example: For elacestrant (ER degrader), comparators = fulvestrant, other SERDs
2. For each comparator, run abbreviated tool chain:
a. PubChem_get_CID_by_compound_name(compound=comparator)
b. ChEMBL_search_activities(chemblid=comparator_chemblid, target="ESR1", max_results=20)
└─ Extract: Potency vs primary target
c. search_clinical_trials(intervention=comparator, condition=indication, pageSize=20)
└─ Extract: Phase 3 trial counts, approval status
d. FAERS_count_reactions_by_drug_event(medicinalproduct=comparator)
└─ Extract: Top 5 adverse events, seriousness ratio
3. Search for head-to-head trials:
search_clinical_trials(intervention=f"{drug_name} AND {comparator}")
└─ Extract: Direct comparison trials [★★★]
4. PubMed_search_articles(query=f"{drug_name} vs {comparator}", max_results=10)
└─ Extract: Network meta-analyses, indirect comparisons [★★☆]
5. Create comparison tables across dimensions:
└─ Potency, selectivity, ADMET, efficacy, safety, cost (if available)
Output for Section 10.5:
### 10.5 Comparative Analysis
#### Drug Class: Selective Estrogen Receptor Degraders (SERDs)
**Primary Comparators**: Fulvestrant (approved), AZD9833 (investigational), GDC-9545 (investigational)
#### Potency Comparison
| Drug | ESR1 WT IC50 | ESR1 Y537S IC50 | Selectivity | Source |
|------|-------------|-----------------|-------------|--------|
| **Elacestrant** | 48 nM | 77 nM | > 100x vs other NRs | ChEMBL |
| Fulvestrant | 9 nM | ~50 nM (est) | > 100x | ChEMBL |
| AZD9833 | 0.7 nM | 1.2 nM | > 1000x | Literature |
**Potency Ranking**: AZD9833 > Fulvestrant ≈ Elacestrant for WT; all active against Y537S
*Sources: ChEMBL via `ChEMBL_search_activities`, PMID:33445678*
#### Clinical Trial Landscape
| Drug | Phase 3 Trials | Primary Indication | Approval Status |
|------|----------------|-------------------|-----------------|
| Elacestrant | 2 completed, 1 ongoing | ER+/HER2- mBC | Approved (US, 2023) |
| Fulvestrant | 15+ completed | ER+/HER2- mBC | Approved (2002) |
| AZD9833 | 3 ongoing | ER+/HER2- mBC | Investigational |
| GDC-9545 | 2 ongoing | ER+/HER2- mBC | Investigational |
*Source: ClinicalTrials.gov*
#### Safety Profile Comparison
| Drug | Top AE (% patients) | Serious AE Rate | Fatal Outcomes |
|------|---------------------|-----------------|----------------|
| Elacestrant | Nausea (35%), Fatigue (30%) | 51.8% | 4.7% (FAERS) |
| Fulvestrant | Injection site reaction (40%), Hot flash (28%) | 48.2% | 3.9% (FAERS) |
**Safety Differentiation**: Elacestrant oral administration avoids injection site reactions but has higher GI AE rate.
*Sources: FAERS via `FAERS_count_reactions_by_drug_event`, product labels*
#### Head-to-Head Trials
**EMERALD vs Fulvestrant**:
- Trial: NCT03778931 (Phase 3, N=478, completed)
- PFS: 3.8 mo (elacestrant) vs 1.9 mo (fulvestrant) in ESR1-mutated subgroup (HR 0.55, p<0.001)
- PFS: 2.2 mo vs 1.9 mo in overall population (HR 0.84, p=0.05)
*Source: `extract_clinical_trial_outcomes` (NCT03778931)*
#### Differentiation Factors
| Factor | Elacestrant Advantage | Fulvestrant Advantage |
|--------|----------------------|----------------------|
| **Route** | Oral (QD) | IM injection (Q4W after loading) |
| **ESR1 mutant efficacy** | +100% PFS improvement | Less data |
| **Brain metastases** | BBB penetration (preclinical) | Poor CNS penetration |
| **Approval** | Biomarker-selected (ESR1 mut) | Broader indication |
| **Experience** | Limited (1 yr post-approval) | Extensive (20+ yrs) |
**Positioning**: Elacestrant fills unmet need for oral SERD with superior efficacy in ESR1-mutated disease. Fulvestrant remains standard for ESR1 WT due to longer track record.
Type Normalization & Error Prevention
Common Validation Errors
Many ToolUniverse tools require string inputs but may return integers or floats. Always convert IDs to strings.
Problem Examples:
- ChEMBL target IDs:
12345(int) → should be"12345"(str) - PubMed IDs:
23456789(int) → should be"23456789"(str) - Clinical trial NCT IDs: sometimes parsed as numbers
Type Normalization Helper
Before calling any tool with ID parameters:
# Convert all IDs to strings
chembl_ids = [str(id) for id in chembl_ids]
nct_ids = [str(id) for id in nct_ids]
pmids = [str(id) for id in pmids]
Pre-Call Checklist
Before each API call:
- All ID parameters are strings
- Lists contain strings, not ints/floats
- No
Noneornullvalues in required fields - Arrays are non-empty if required
Evidence Grading System
Evidence Tiers
| Tier | Symbol | Description | Example |
|---|---|---|---|
| T1 | ★★★ | Phase 3 RCT, meta-analysis, FDA approval | Pivotal trial, label indication |
| T2 | ★★☆ | Phase 1/2 trial, large case series | Dose-finding study |
| T3 | ★☆☆ | In vivo animal, in vitro cellular | Mouse PK study |
| T4 | ☆☆☆ | Review mention, computational prediction | ADMET-AI prediction |
Application in Report
Metformin reduces hepatic glucose output via AMPK activation [★★★: FDA Label].
Phase 3 trials demonstrated HbA1c reduction of 1.0-1.5% [★★★: NCT00123456].
Preclinical studies suggest anti-cancer properties [★☆☆: PMID:23456789].
ADMET-AI predicts low hERG liability (0.12) [☆☆☆: computational].
Per-Section Summary
Include evidence quality summary for each major section:
### 5. Clinical Development
**Evidence Quality**: Strong (156 Phase 3 trials, 203 Phase 2, 67 Phase 1)
**Data Confidence**: High - mature clinical program with decades of data
Section Completeness Checklist
Before finalizing any report, verify each section meets minimum requirements:
Section 1 (Identity) - Minimum Requirements
- PubChem CID with link
- ChEMBL ID with link (or "Not in ChEMBL")
- Canonical SMILES
- Molecular formula and weight
- At least 3 brand names OR "Generic only"
- Salt forms identified (or "Parent compound only")
Section 2 (Chemistry) - Minimum Requirements
- 6+ physicochemical properties in table format (including pKa if available)
- Lipinski rule assessment with pass/fail
- QED score with interpretation
- Solubility data (predicted or label-based)
- Salt forms documented (or "Parent compound only")
- 2D structure image embedded (PubChem link)
- Formulation details if available (dosage forms, excipients)
Section 3 (Mechanism) - Minimum Requirements
- FDA label MOA text quoted (if approved drug) OR literature MOA summary
- Primary mechanism described in 2-3 sentences
- At least 1 primary target with UniProt ID
- Activity type and potency (IC50/EC50/Ki) with assay count
- Target selectivity table (including mutant forms if relevant, e.g., ESR1 Y537S for endocrine drugs)
- Off-target activity addressed (or "Highly selective")
Section 4 (ADMET) - Minimum Requirements
- All 5 subsections present (A, D, M, E, T)
- Absorption: bioavailability + at least 2 other endpoints (predicted OR label PK)
- Distribution: BBB + VDss or PPB (predicted OR label PK)
- Metabolism: CYP substrate/inhibitor status for 3+ CYPs (predicted OR label DDI)
- Excretion: clearance OR half-life (predicted OR label PK)
- Toxicity: AMES + hERG + at least 1 other (predicted OR label warnings)
- If ADMET-AI fails, fallback to FDA label PK sections (do NOT leave "predictions unavailable")
Section 5 (Clinical) - Minimum Requirements
- Development status clearly stated (Approved/Investigational/Preclinical)
- Actual counts by phase/status in table format (NOT just representative trial list)
- Indication breakdown by counts (e.g., "312 diabetes trials, 87 PCOS trials")
- Approved indications with year (or "Not approved")
- Representative trial list (top 5 Phase 3, top 3 recruiting) with clear labels
- Key efficacy data with trial references (or "No outcome data available")
Section 6 (Safety) - Minimum Requirements
- Top 5 adverse events with frequencies
- FAERS seriousness breakdown (serious vs non-serious counts)
- FAERS date window documented (e.g., "2004-2026")
- FAERS limitations paragraph (small N, reporting bias, causality not established)
- Black box warnings (or "None")
- At least 3 drug-drug interactions with mechanism (CYP, transporter) OR "No significant interactions"
- Dose modification triggers (ALT/AST thresholds, renal impairment, CYP inhibitor/inducer adjustments)
Section 7 (PGx) - Minimum Requirements
- Pharmacogenes listed (or "None identified")
- CPIC/DPWG guideline status (or "No guideline available")
- At least 1 clinical annotation OR "No annotations identified"
- If PharmGKB fails, fallback to label PGx sections + literature (document the failure)
Section 10 (Conclusions) - Minimum Requirements
- 5-point scorecard covering: efficacy, safety, PK, druggability, competition
- 3+ key strengths
- 3+ key concerns/limitations
- At least 2 research gaps identified
Drug Profile Scorecard Template
Include in Section 10:
### 10.1 Drug Profile Scorecard
| Criterion | Score (1-5) | Rationale |
|-----------|-------------|-----------|
| **Efficacy Evidence** | 5 | Multiple Phase 3 trials, decades of use |
| **Safety Profile** | 4 | Well-tolerated; lactic acidosis rare but serious |
| **PK/ADMET** | 4 | Good bioavailability; renal elimination |
| **Target Validation** | 4 | AMPK mechanism well-established |
| **Competitive Position** | 3 | First-line but many alternatives |
| **Overall** | 4.0 | **Strong drug profile** |
**Interpretation**:
- 5 = Excellent, 4 = Good, 3 = Moderate, 2 = Concerning, 1 = Poor
Automated Completeness Audit
CRITICAL: Before finalizing the report, run this audit checklist and append findings to Section 11.
Audit Process
- Review each section against minimum requirements (see Section Completeness Checklist)
- Flag any missing data with specific tool call recommendations
- Document tool failures and fallback attempts
- Generate completeness score (% of minimum requirements met)
Audit Output Template
Add this to Section 11 (Data Sources & Methodology):
---
## Report Completeness Audit
**Overall Completeness**: 85% (17/20 minimum requirements met)
### Missing Data Items
| Section | Missing Item | Recommended Action |
|---------|--------------|-------------------|
| 2 | Salt forms | Call `DailyMed_get_spl_sections_by_setid` (chemistry section) |
| 3 | Mutant ESR1 binding | Filter ChEMBL activities for ESR1 Y537S, D538G variants |
| 5 | Phase count breakdown | Compute counts from `search_clinical_trials` results |
| 7 | PharmGKB guidelines | PharmGKB API unavailable; used label PGx instead [✓] |
### Tool Failures Encountered
| Tool | Error | Fallback Used |
|------|-------|---------------|
| `PharmGKB_search_drugs` | API timeout | DailyMed label PGx sections [✓] |
| `ADMETAI_predict_toxicity` | Invalid SMILES | FDA label warnings section [✓] |
### Data Confidence Assessment
| Section | Confidence | Evidence Tier | Notes |
|---------|-----------|---------------|-------|
| 1. Identity | High | ★★★ | PubChem + ChEMBL confirmed |
| 2. Chemistry | Medium | ★★☆ | Missing salt form details |
| 3. Mechanism | High | ★★★ | FDA label + ChEMBL bioactivity |
| 4. ADMET | Medium | ★★☆ | Predictions only; no clinical PK |
| 5. Clinical | High | ★★★ | 156 Phase 3 trials analyzed |
| 6. Safety | High | ★★★ | FAERS + label warnings |
| 7. PGx | Low | ★☆☆ | PharmGKB unavailable; label only |
### Quality Control Metrics (Section 11.3)
#### Data Recency
| Source | Last Updated | Data Age | Status |
|--------|-------------|----------|--------|
| PubChem | 2026-02-01 | < 1 week | ✓ Current |
| ChEMBL v33 | 2025-12-15 | 2 months | ✓ Current |
| FAERS | 2026-01-01 (2026Q1) | < 1 month | ✓ Current |
| DailyMed | 2025-11-20 (label revised) | 3 months | ✓ Current |
| PharmGKB | N/A (unavailable) | - | ⚠ Missing |
**Recency Assessment**: All data sources current (< 6 months). PharmGKB unavailable; fallback used.
#### Cross-Source Validation
| Property | PubChem | ChEMBL | DailyMed | Agreement |
|----------|---------|--------|----------|-----------|
| Molecular Weight | 378.88 | 378.88 | 378.88 | ✓ Exact match |
| Half-life | N/A | N/A | 27 hours | Single source |
| Primary target | N/A | ESR1 | ESR1 | ✓ Confirmed |
| Bioavailability | Predicted: 85% | N/A | ~60% (fed) | ⚠ Discrepancy |
**Contradictions Detected**:
- Bioavailability: ADMET-AI predicts 85%, but label reports ~60% (fed state). **Resolution**: Use label value (T1: ★★★) over prediction (T2: ★★☆).
#### Completeness Score
**Overall**: 85% (17/20 minimum requirements met)
| Category | Score | Details |
|----------|-------|---------|
| Identity & Structure | 100% | 5/5 - All identifiers present |
| Chemistry | 80% | 4/5 - Missing salt form |
| Mechanism | 90% | 9/10 - Minor gap in off-targets |
| Clinical Development | 95% | 19/20 - Comprehensive trial data |
| Safety | 100% | 10/10 - FAERS + label complete |
| Pharmacogenomics | 60% | 3/5 - PharmGKB unavailable |
| Regulatory | 80% | 4/5 - US only, no EMA/PMDA |
#### Evidence Distribution
| Tier | Count | Percentage | Interpretation |
|------|-------|------------|----------------|
| T1 (★★★) | 45 | 65% | High-quality regulatory/experimental |
| T2 (★★☆) | 18 | 26% | Computational predictions, PharmGKB |
| T3 (★☆☆) | 5 | 7% | Literature inference |
| T4 (☆☆☆) | 1 | 1% | Speculation |
**Quality Assessment**: 91% of claims backed by T1/T2 evidence. Report meets publication standards.
**Recommendation**: Address missing items in Sections 2, 3, 5 for publication-quality report.
Fallback Chains
| Primary Tool | Fallback | Use When |
|---|---|---|
PubChem_get_CID_by_compound_name |
ChEMBL_search_compounds |
Name not in PubChem |
ChEMBL_get_molecule_targets |
Use ChEMBL_search_activities instead |
Avoid this tool (returns irrelevant targets) |
ChEMBL_get_bioactivity_by_chemblid |
PubChem_get_bioactivity_summary_by_CID |
No ChEMBL ID |
DailyMed_search_spls |
PubChem_get_drug_label_info_by_CID |
DailyMed timeout |
PharmGKB_get_dosing_guidelines |
DailyMed_get_spl_sections_by_setid (pharmacogenomics) |
PharmGKB API error |
PharmGKB_search_drugs |
DailyMed_get_spl_sections_by_setid + PubMed_search_articles |
PharmGKB unavailable |
FAERS_count_reactions_by_drug_event |
Document "FAERS unavailable" + use label AEs | API error |
ADMETAI_* (all tools) |
DailyMed_get_spl_sections_by_setid (clinical_pharmacology, pharmacokinetics) |
Invalid SMILES or API error |
Quick Reference: Tools by Use Case
| Use Case | Primary Tool | Fallback | Evidence |
|---|---|---|---|
| Name → CID | PubChem_get_CID_by_compound_name |
ChEMBL_search_compounds |
★★★ |
| SMILES → CID | PubChem_get_CID_by_SMILES |
- | ★★★ |
| Properties | PubChem_get_compound_properties_by_CID |
ADMETAI_predict_physicochemical_properties |
★★★ / ★★☆ |
| Salt forms | DailyMed_get_spl_sections_by_setid (chemistry) |
- | ★★★ |
| Formulation | DailyMed_get_spl_sections_by_setid (description, inactive_ingredients) |
- | ★★★ |
| Drug-likeness | ADMETAI_predict_physicochemical_properties |
Calculate from properties | ★★☆ |
| FDA MOA | DailyMed_get_spl_sections_by_setid (mechanism_of_action) |
- | ★★★ |
| Targets | ChEMBL_search_activities → ChEMBL_get_target |
DGIdb_get_drug_info |
★★★ |
| Avoid | ChEMBL_get_molecule_targets |
Use activities-based approach | N/A |
| Bioactivity | ChEMBL_search_activities |
PubChem_get_bioactivity_summary_by_CID |
★★★ |
| Absorption | ADMETAI_predict_bioavailability |
DailyMed clinical_pharmacology |
★★☆ / ★★★ |
| BBB | ADMETAI_predict_BBB_penetrance |
DailyMed clinical_pharmacology |
★★☆ / ★★★ |
| CYP | ADMETAI_predict_CYP_interactions |
DailyMed drug_interactions |
★★☆ / ★★★ |
| Toxicity | ADMETAI_predict_toxicity |
DailyMed warnings_and_cautions |
★★☆ / ★★★ |
| Trials | search_clinical_trials |
- | ★★★ |
| Phase counts | Compute from search_clinical_trials results |
- | ★★★ |
| Trial outcomes | extract_clinical_trial_outcomes |
- | ★★★ |
| FAERS | FAERS_count_reactions_by_drug_event |
Label adverse_reactions | ★★★ |
| Dose mods | DailyMed_get_spl_sections_by_setid (dosage_and_administration, warnings) |
- | ★★★ |
| Label | DailyMed_search_spls |
PubChem_get_drug_label_info_by_CID |
★★★ |
| PGx | PharmGKB_search_drugs |
DailyMed pharmacogenomics + PubMed |
★★☆ / ★★★ |
| CPIC | PharmGKB_get_dosing_guidelines |
DailyMed pharmacogenomics |
★★★ / ★★☆ |
| Literature | PubMed_search_articles |
EuropePMC_search_articles |
Varies |
Common Use Cases
Approved Drug Profile
User: "Tell me about metformin" → Full 11-section report emphasizing clinical data, FAERS, PGx
Investigational Compound
User: "What do we know about compound X (ChEMBL123456)?" → Emphasize preclinical data, mechanism, early trials; safety sections may be sparse
Safety Review
User: "What are the safety concerns with drug Y?" → Deep dive on FAERS, black box warnings, interactions, PGx; lighter on chemistry
ADMET Assessment
User: "Evaluate this compound's drug-likeness [SMILES]" → Focus on Sections 2 and 4; other sections may be brief or N/A
Clinical Development Landscape
User: "What trials are ongoing for drug Z?" → Heavy emphasis on Section 5; trial tables with status, phases, indications
When NOT to Use This Skill
- Target research → Use target-intelligence-gatherer skill
- Disease research → Use disease-research skill
- Literature-only → Use literature-deep-research skill
- Single property lookup → Call tool directly
- Structure similarity search → Use
PubChem_search_compounds_by_similaritydirectly
Use this skill for comprehensive, multi-dimensional drug profiling.
Key Improvements Summary
Based on real-world testing (elacestrant case study), these workflow improvements address common gaps:
1. Chemistry Completeness
- ✅ Add salt/polymorph information from DailyMed chemistry section
- ✅ Include pKa and experimental solubility
- ✅ Embed 2D structure image (PubChem link)
- ✅ Document formulation details and excipients
2. Mechanism Depth
- ✅ Quote FDA label MOA text verbatim (authoritative source)
- ✅ Add target selectivity table (including mutant forms for relevant drugs)
- ✅ Derive targets from ChEMBL activities (avoid
ChEMBL_get_molecule_targets)
3. Clinical Trial Accuracy
- ✅ Compute actual phase counts from search results
- ✅ Separate by indication when relevant (e.g., breast cancer vs other)
- ✅ Clearly label "representative trials" vs "all trials"
4. Safety Completeness
- ✅ Add FAERS seriousness breakdown
- ✅ Document date window for FAERS data
- ✅ Include FAERS limitations paragraph
- ✅ Add dose modification triggers table (ALT/AST thresholds, renal/hepatic dosing)
5. Fallback Resilience
- ✅ ADMET-AI failures → FDA label PK sections
- ✅ PharmGKB failures → DailyMed PGx sections + PubMed
- ✅ Type normalization for all IDs (prevent validation errors)
6. Quality Control
- ✅ Automated completeness audit at report end
- ✅ Missing data flagged with specific tool call recommendations
- ✅ Tool failures documented with fallback status
Additional Resources
- Tool reference: TOOLS_REFERENCE.md - Complete tool listing
- Verification checklist: CHECKLIST.md - Pre-delivery verification
- Examples: EXAMPLES.md - Detailed workflow examples